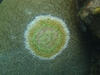
We investigated coral disease processes and causes by characterizing microbial communities in diseased and healthy representatives of selected coral species both temporally and spatially by employing microarray technology. We tested the diagnostic potential of coral fluorescence for identifying disease-induced physiological stress.
Coral disease
Coral diseases were first reported on reefs in the Florida Keys and Caribbean in the 1970s. In the decades since, they have been reported worldwide and with increasing frequency. Disease is now recognized as one of the major causes of reef degradation and coral mortality. Recent research has suggested that coral diseases may be secondary opportunistic infections, rather than the result of primary pathogens, making it imperative to understand the microbial shifts that accompany the transition from healthy to diseased corals. Additionally, we need to determine if the spread of coral disease is affected by the level of connectivity among water masses, organisms, trophic levels, or habitats. See black-band disease and coral bleaching galleries below.
We investigated coral disease processes and causes by characterizing microbial communities in diseased and healthy representatives of selected coral species both temporally and spatially by employing microarray technology. We tested the diagnostic potential of coral fluorescence for identifying disease-induced physiological stress. This work links coral ecosystem studies in marine protected areas to better understand coral ecosystem health.
Specific research efforts included:
-
Comparing microbial communities between diseased and healthy corals from two species in two National Parks: Dry Tortugas National Park and Virgin Islands National Park.
-
Utilizing microarray technology (representing 30,000 microbial taxa) to get a taxonomic overview of the shifts in microbial communities between healthy and diseased corals, between species of corals, and between different geographic areas
-
Comparison of DNA preservation methods for environmental bacterial community samples.
Field collections of environmental samples, for example corals, for molecular microbial analyses present distinct challenges. The lack of laboratory facilities in remote locations is common, and preservation of microbial community DNA for later study is critical. A particular challenge is keeping samples frozen in transit.
Five preservation methods that do not require cold storage were compared for effectiveness over time and ease of use. Mixed microbial communities of known composition were created and preserved by DNAgard™, RNAlater®, DMSO–EDTA–salt (DESS), FTA® cards, and FTA Elute® cards. Microbial community fingerprinting analysis and DNA sequencing were used to detect specific changes in the known communities over weeks and months of storage. A previously known bias in FTA® cards that results in lower recovery of pure cultures of Gram-positive bacteria was also detected in mixed community samples. There appears to be a uniform bias across all five preservation methods against microorganisms with high G + C DNA. Overall, the liquid-based preservatives (DNAgard™, RNAlater®, and DESS) outperformed the card-based methods. No single liquid method clearly outperformed the others, leaving method choice to be based on experimental design, field facilities, shipping constraints, and allowable cost.
-
Field testing a new technique to determine if coral fluorescence can be used to remotely assess coral health
Note: Any use of trade, product, or firm names is for descriptive purposes only and does not imply endorsement by the U.S. Government.
Below are other science projects associated with this project.
Coral Reef Ecosystem Studies (CREST)
Applications of Coral Fluorescence
Below are publications associated with this project.
Comparing bacterial community composition of healthy and dark spot-affected Siderastrea siderea in Florida and the Caribbean Comparing bacterial community composition of healthy and dark spot-affected Siderastrea siderea in Florida and the Caribbean
Comparison of three DNA extraction kits to establish maximum yield and quality of coral-associated microbial DNA Comparison of three DNA extraction kits to establish maximum yield and quality of coral-associated microbial DNA
Evaluation of coral pathogen growth rates after exposure to atmospheric African dust samples Evaluation of coral pathogen growth rates after exposure to atmospheric African dust samples
Comparing bacterial community composition between healthy and white plague-like disease states in Orbicella annularis using PhyloChip™ G3 microarrays Comparing bacterial community composition between healthy and white plague-like disease states in Orbicella annularis using PhyloChip™ G3 microarrays
Comparison of DNA preservation methods for environmental bacterial community samples Comparison of DNA preservation methods for environmental bacterial community samples
PhyloChipTM microarray comparison of sampling methods used for coral microbial ecology PhyloChipTM microarray comparison of sampling methods used for coral microbial ecology
Microbial ecology of corals, sponges, and algae in mesophotic coral environments Microbial ecology of corals, sponges, and algae in mesophotic coral environments
Applying New Methods to Diagnose Coral Diseases Applying New Methods to Diagnose Coral Diseases
Cross-kingdom amplification using Bacteria-specific primers: Complications for studies of coral microbial ecology Cross-kingdom amplification using Bacteria-specific primers: Complications for studies of coral microbial ecology
We investigated coral disease processes and causes by characterizing microbial communities in diseased and healthy representatives of selected coral species both temporally and spatially by employing microarray technology. We tested the diagnostic potential of coral fluorescence for identifying disease-induced physiological stress.
Coral disease
Coral diseases were first reported on reefs in the Florida Keys and Caribbean in the 1970s. In the decades since, they have been reported worldwide and with increasing frequency. Disease is now recognized as one of the major causes of reef degradation and coral mortality. Recent research has suggested that coral diseases may be secondary opportunistic infections, rather than the result of primary pathogens, making it imperative to understand the microbial shifts that accompany the transition from healthy to diseased corals. Additionally, we need to determine if the spread of coral disease is affected by the level of connectivity among water masses, organisms, trophic levels, or habitats. See black-band disease and coral bleaching galleries below.
We investigated coral disease processes and causes by characterizing microbial communities in diseased and healthy representatives of selected coral species both temporally and spatially by employing microarray technology. We tested the diagnostic potential of coral fluorescence for identifying disease-induced physiological stress. This work links coral ecosystem studies in marine protected areas to better understand coral ecosystem health.
Specific research efforts included:
-
Comparing microbial communities between diseased and healthy corals from two species in two National Parks: Dry Tortugas National Park and Virgin Islands National Park.
-
Utilizing microarray technology (representing 30,000 microbial taxa) to get a taxonomic overview of the shifts in microbial communities between healthy and diseased corals, between species of corals, and between different geographic areas
-
Comparison of DNA preservation methods for environmental bacterial community samples.
Field collections of environmental samples, for example corals, for molecular microbial analyses present distinct challenges. The lack of laboratory facilities in remote locations is common, and preservation of microbial community DNA for later study is critical. A particular challenge is keeping samples frozen in transit.
Five preservation methods that do not require cold storage were compared for effectiveness over time and ease of use. Mixed microbial communities of known composition were created and preserved by DNAgard™, RNAlater®, DMSO–EDTA–salt (DESS), FTA® cards, and FTA Elute® cards. Microbial community fingerprinting analysis and DNA sequencing were used to detect specific changes in the known communities over weeks and months of storage. A previously known bias in FTA® cards that results in lower recovery of pure cultures of Gram-positive bacteria was also detected in mixed community samples. There appears to be a uniform bias across all five preservation methods against microorganisms with high G + C DNA. Overall, the liquid-based preservatives (DNAgard™, RNAlater®, and DESS) outperformed the card-based methods. No single liquid method clearly outperformed the others, leaving method choice to be based on experimental design, field facilities, shipping constraints, and allowable cost.
-
Field testing a new technique to determine if coral fluorescence can be used to remotely assess coral health
Note: Any use of trade, product, or firm names is for descriptive purposes only and does not imply endorsement by the U.S. Government.
Below are other science projects associated with this project.
Coral Reef Ecosystem Studies (CREST)
Applications of Coral Fluorescence
Below are publications associated with this project.